Structural insight into D -xylose utilization by xylose reductase from Scheffersomyces stipitis | Scientific Reports

L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study - ScienceDirect

d‐Xylose consumption by nonrecombinant Saccharomyces cerevisiae: A review - Patiño - 2019 - Yeast - Wiley Online Library
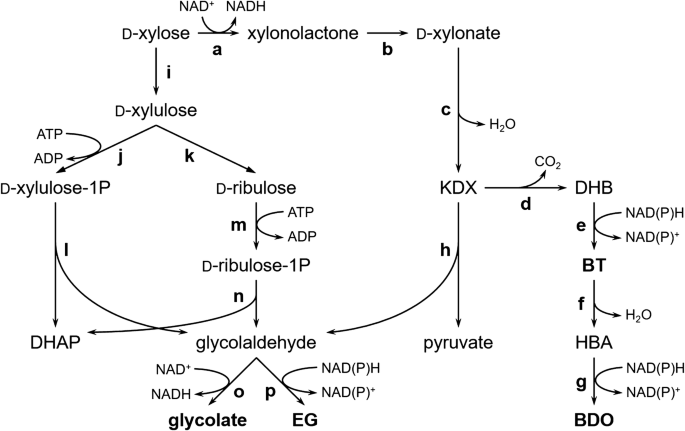
Biochemical routes for uptake and conversion of xylose by microorganisms | Biotechnology for Biofuels | Full Text

Structural insight into D -xylose utilization by xylose reductase from Scheffersomyces stipitis | Scientific Reports

Synthetic (blue) and natural (black) d-xylose assimilation pathways. In... | Download Scientific Diagram

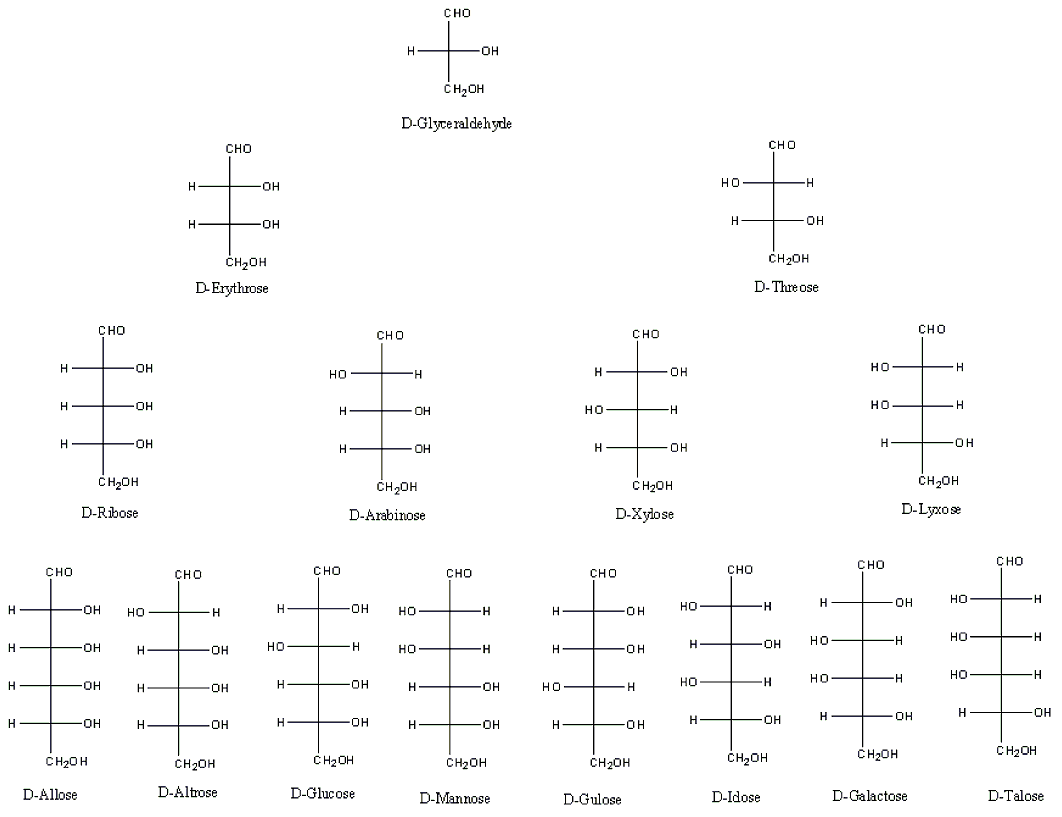
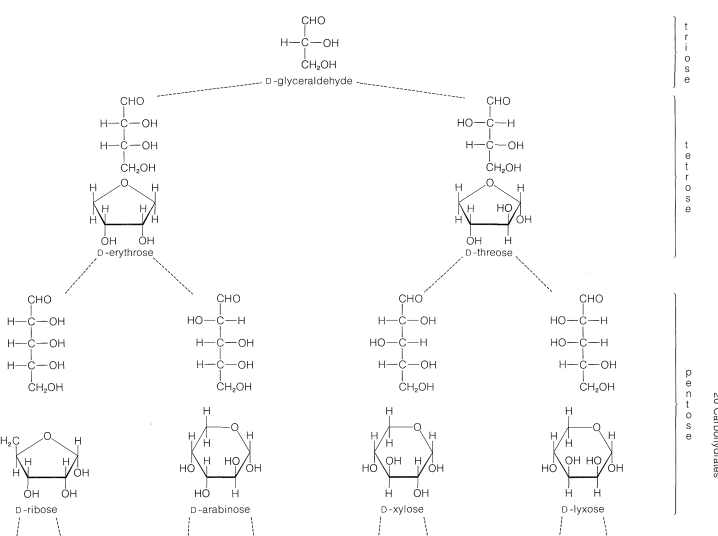
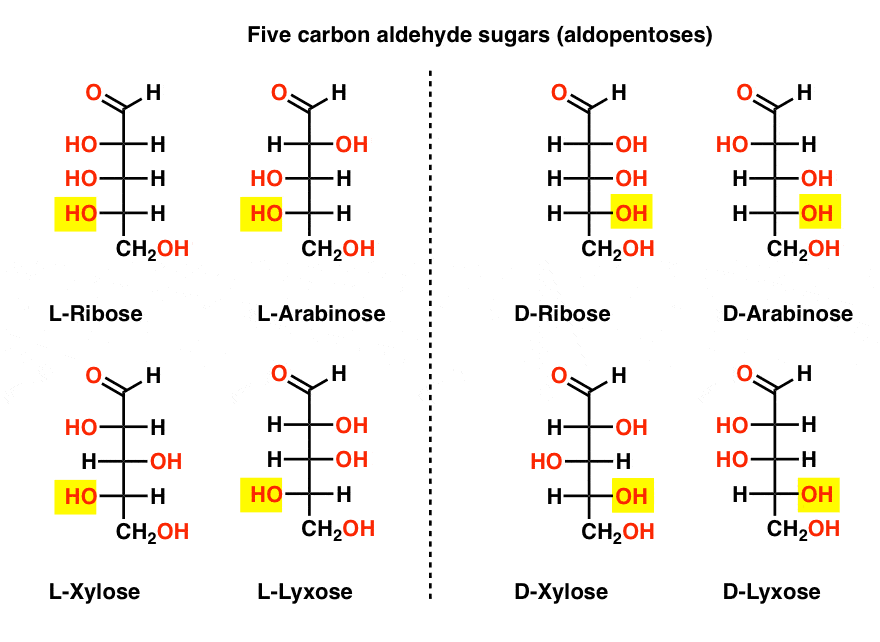
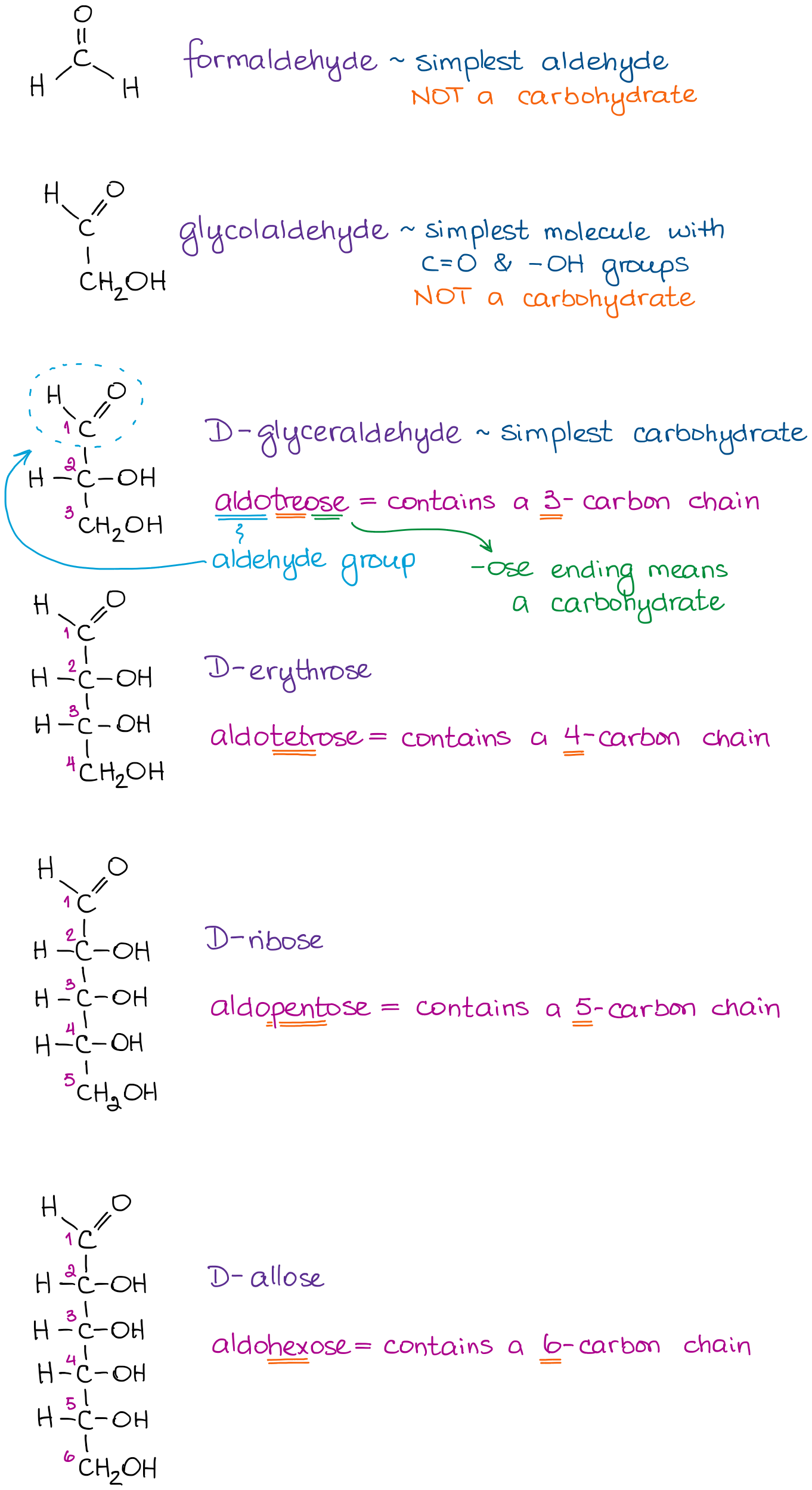
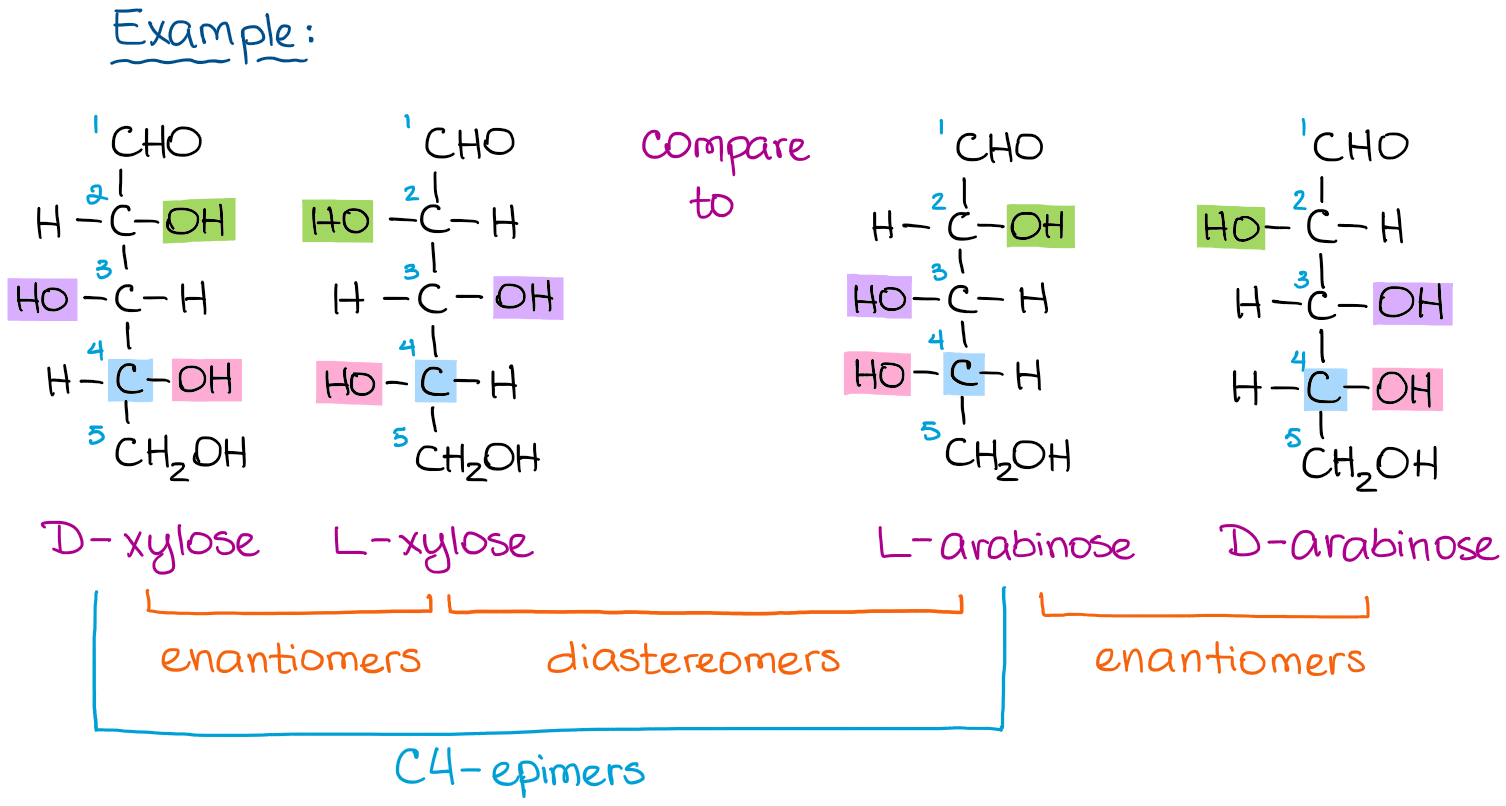
Chemical structures of aldopentose stereoisomers (Fischer projection).... | Download Scientific Diagram
![PDF] Observations of reaction intermediates and the mechanism of aldose-ketose interconversion by D-xylose isomerase. | Semantic Scholar PDF] Observations of reaction intermediates and the mechanism of aldose-ketose interconversion by D-xylose isomerase. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/02d61692f222aa734696b1b45192ff03e7e9f61b/2-Figure1-1.png)
PDF] Observations of reaction intermediates and the mechanism of aldose-ketose interconversion by D-xylose isomerase. | Semantic Scholar