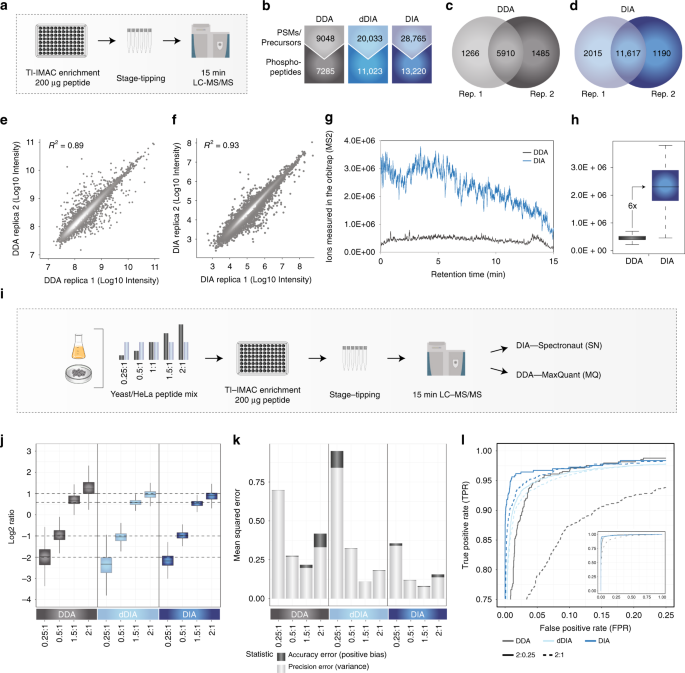
Optimization of Experimental Parameters in Data-Independent Mass Spectrometry Significantly Increases Depth and Reproducibility

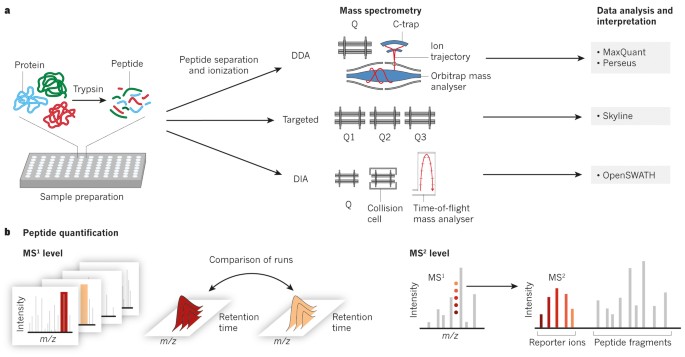
Data‐independent acquisition‐based SWATH‐MS for quantitative proteomics: a tutorial | Molecular Systems Biology
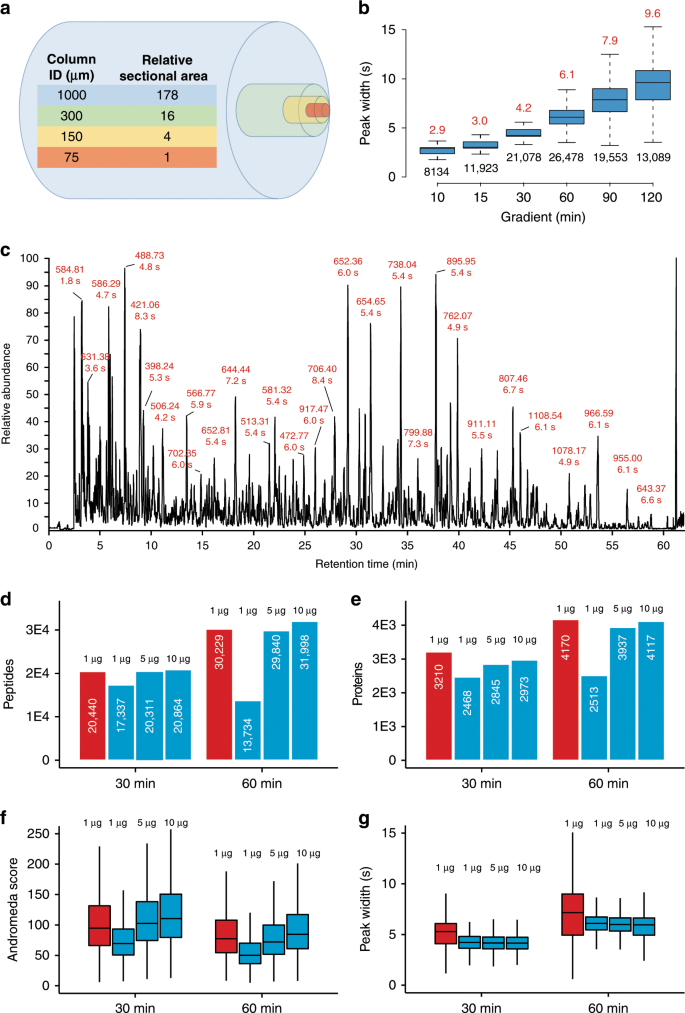
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS | Nature Communications

Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry | Nature Methods

Machine Learning in Mass Spectrometric Analysis of DIA Data - Xu - - PROTEOMICS - Wiley Online Library
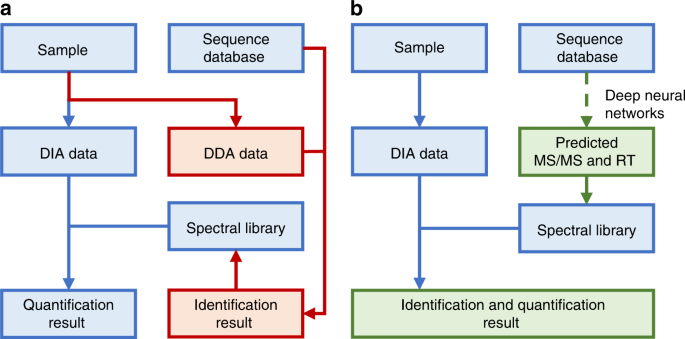
In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics | Nature Communications

The Human Immunopeptidome Project: A Roadmap to Predict and Treat Immune Diseases | Molecular & Cellular Proteomics
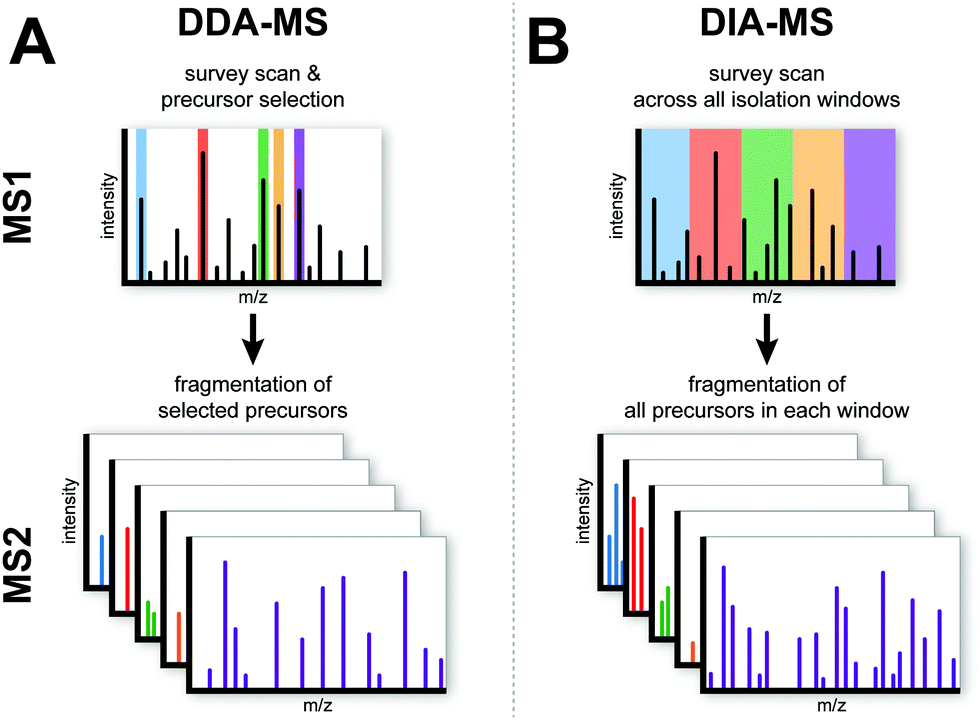
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
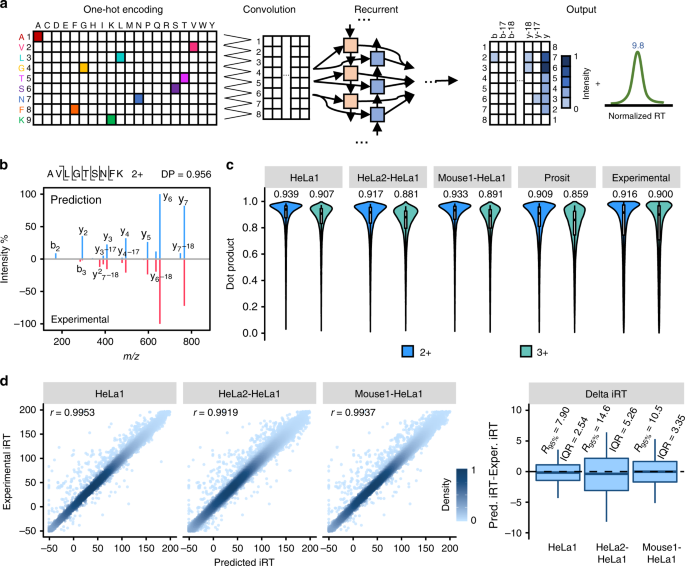
In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics | Nature Communications
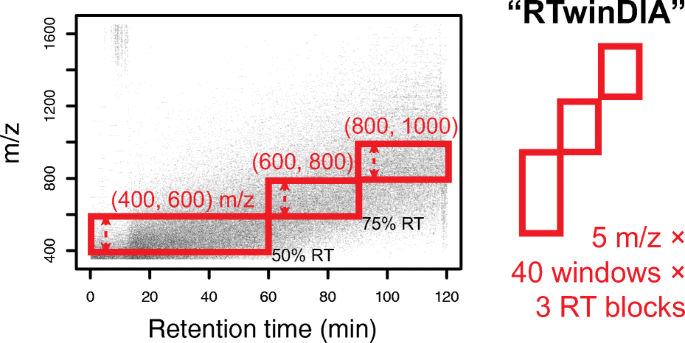
Assessing the Relationship Between Mass Window Width and Retention Time Scheduling on Protein Coverage for Data-Independent Acquisition | SpringerLink

Hybrid Spectral Library Combining DIA-MS Data and a Targeted Virtual Library Substantially Deepens the Proteome Coverage - ScienceDirect

Discovering cellular protein‐protein interactions: Technological strategies and opportunities - Titeca - 2019 - Mass Spectrometry Reviews - Wiley Online Library

Data‐independent acquisition‐based SWATH‐MS for quantitative proteomics: a tutorial | Molecular Systems Biology

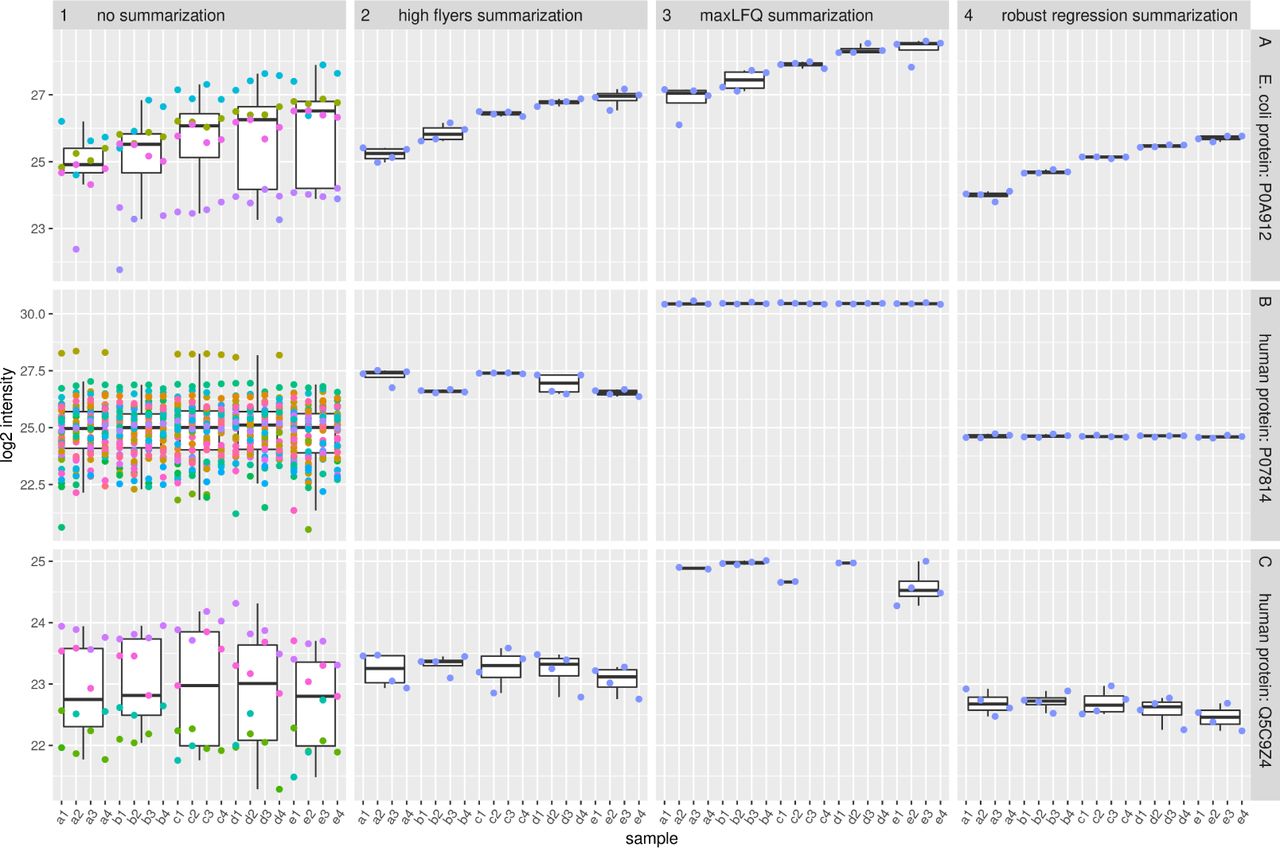
Simultaneous Improvement in the Precision, Accuracy, and Robustness of Label-free Proteome Quantification by Optimizing Data Manipulation Chains | Molecular & Cellular Proteomics
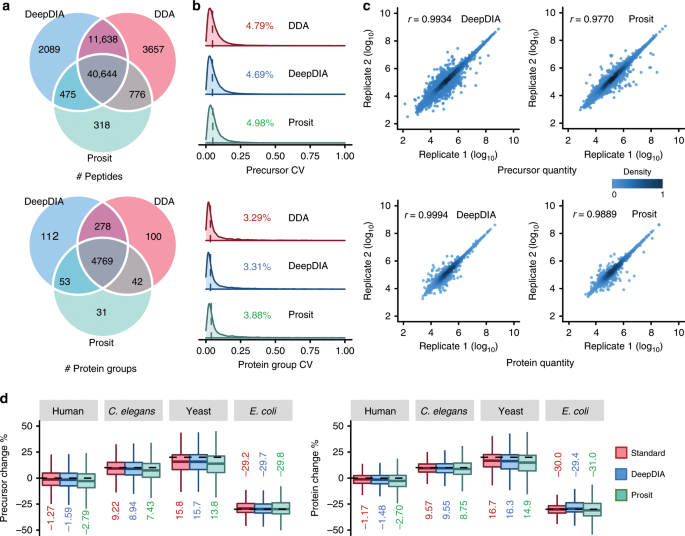
In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics | Nature Communications

PulseDIA: in-depth data independent acquisition mass spectrometry using enhanced gas phase fractionation | bioRxiv